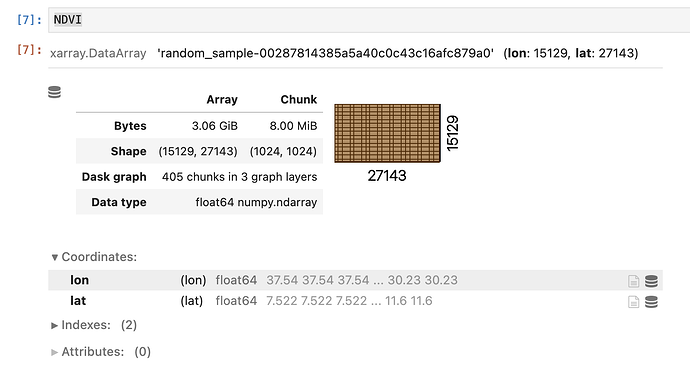
I need to compute and assign the 90th percentile of NDVI values grouped accordingly to some ecoregions. To be more clear, assignment has to replace original NDVI values with the corresponded percentiles based on their associated ecoregion.
Bottle neck is the quantile computation, I was not able to find the way to compute in parallel. Is there any other strategy to make this more efficient?
Assignment with delayed approach fails as well but would be nice to achieve that part in parallel as well.
Data
x, y = [27143, 15129]
# Create dummy data for the coordinates
lat = np.linspace(7.522, 11.6, x)
lon = np.linspace(30.23, 37.54, y)[::-1]
# Create dummy data for the ecoregions
ecoregions = xr.DataArray(
dask.array.random.randint(1, 1000, size=(y, x), chunks=(512, 512)),
coords={
"lon": lon,
"lat": lat,
},
name="ecoregions",
).astype(float)
# Add some nodata values
ecoregions = ecoregions.where(ecoregions < 999)
# Dummy data for the NDVI
NDVI = xr.DataArray(
dask.array.random.random(size=(y, x), chunks=(1024, 1024)),
coords={
"lon": lon,
"lat": lat,
},
)
# Add some nodata values
NDVI = NDVI.where(NDVI < 0.99)
Group and percentile computation
# Statck and rechunk the data
stacked_NDVI = NDVI.stack(z=('lat', 'lon')).chunk({'z': -1})
stacked_ecoregions = ecoregions.stack(z=('lat', 'lon')).chunk({'z': -1})
# Group accordingly by eco-regions and compute the 90th percentile for each of them
quantiles = stacked_NDVI.groupby(stacked_ecoregions).quantile(0.9, skipna=True)
Without stacking and rechunking I was not able to achieve a result. With this small sample it’s really slow to compute it.
Assigment
I’ve tested two options and the eager one is the only working. The other inflate like hell in the ram and then workers die.
Eager :
result = ecoregions.copy()
for ecoregion in quantiles.ecoregions:
result = xr.where(ecoregions == ecoregion, quantiles.sel(ecoregions=ecoregion), result)
Lazy:
from dask import compute, delayed
# Delayed function to assign quantiles
@delayed
def assign_quantiles_to_eco_unit(ecoregions, quantile_values, eco_unit):
return xr.where(ecoregions == eco_unit, quantile_values.sel(eco_units=eco_unit), np.NaN)
# Build the delayed computations
delayed_results = [assign_quantiles_to_eco_unit(ecoregions, quantiles, eco_unit) for eco_unit in quantiles.eco_units]
# Compute the results using Dask
results = compute(*delayed_results)
# Combine the results
final_result = xr.concat(results, dim='dummy').max(dim='dummy', skipna=True)